Software Resources
These software packages were developed by Dan Meliza or by the lab, and are freely available under open-source licenses. Not all code is actively maintained, but we are typically happy to answer questions and to consider pull requests.
Our code for operant hardware and software is described on its own page. All of our publicly available software can be found on the lab github page.
Neuron Models and Data Assimilation
spyks
General-purpose tool to generate fast c++ code for integrating neuron models. Models are specified in a YAML document that contains the equations of motion in symbolic form and values for all of the parameters and state variables. The model may also include a reset rule. Values are specified with physical units for dimensional analysis and reduced errors from unit mismatches.
Hosted on GitHub: melizalab/spyks
Publications:
Chen AN, Meliza CD
(2018). Phasic and Tonic Cell Types in the Zebra Finch Auditory Caudal Mesopallium.
J Neurophys,
doi:10.1152/jn.00694.2017
Bjoring MC, Meliza CD
(2019). A low-threshold potassium current enhances sparseness and reliability in a model of avian auditory cortex,
PLoS Comp Biol,
doi:10.1371/journal.pcbi.1006723
Fehrman C, Robbins TD, Meliza CD
(2021). Nonlinear effects of intrinsic dynamics on temporal encoding in a model of avian auditory cortex,
PLoS Comp Biol,
doi:10.1371/journal.pcbi.1008768
dynamical state and parameter estimation
Example code for dynamical state and parameter estimation of a biophysical neuron model. Given intracellular voltage measurements and a dynamical model, estimates the gating variables, kinetic parameters and conductances of the model.
Hosted on GitHub: melizalab/dpse-example
Publications:
Meliza CD, Kostuk M, Huang H, Nogaret A, Margoliash D, Abarbanel HDI (2014).
Estimating parameters and predicting membrane voltages with
conductance-based neuron models. Biol Cybern,
doi:10.1007/s00422-014-0615-5
Data Acquisition
jill
JILL is a realtime system for auditory behavioral and neuroscience experiments based on the [JACK audio framework](http://jackaudio.org/). It consists of several independent modules that handle stimulus presentation, vocalization detection, and data recording. With JILL, you can:
- record acoustic signals from microphones, triggering acquisition when signals reach a certain loudness
- record extracellular neural data from single or multiple electrodes
- present sets of acoustic stimuli, with control over the order, spacing, and repetition of individual stimuli.
- interface with digital I/O systems for monitoring behavior and giving reinforcement
- link modules in a low-latency real-time framework for closed-loop experiments
- use any existing JACK program, or write your own!
Hosted on GitHub: melizalab/jill
Data and Signal Analysis
quickspikes
Simple threshold-based spike detection, implemented in cython.
Hosted on GitHub: melizalab/quickspikes
chirp
Tools for measuring pitch and comparing vocalizations. Features include:
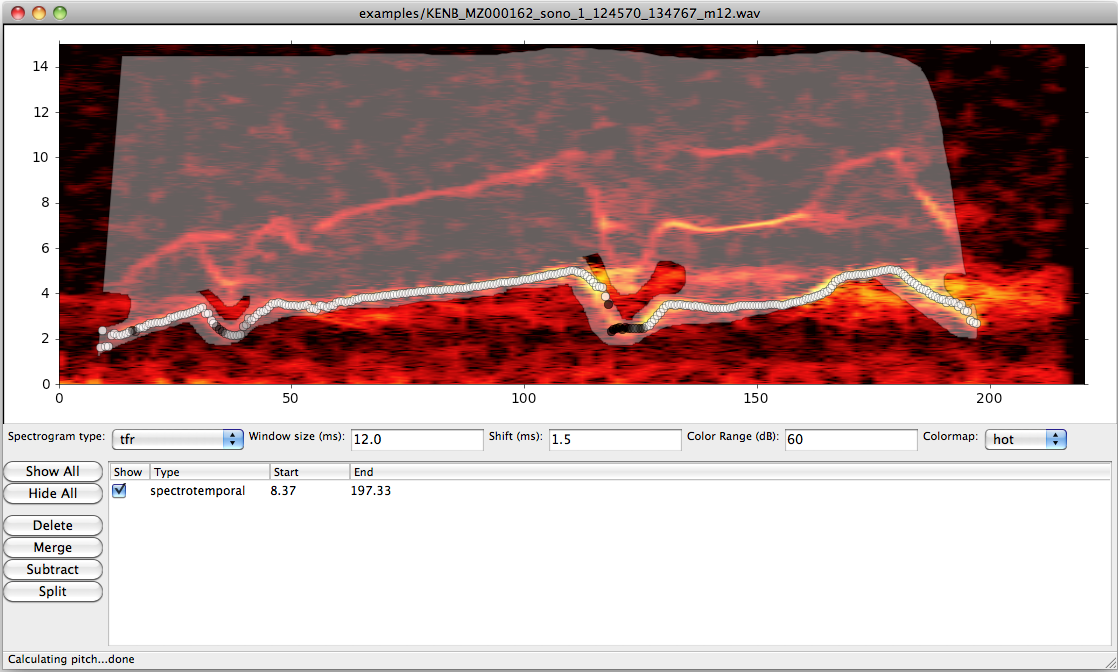
- A graphical interface for examining conventional and time-frequency reassignment spectrograms of recordings, and for labeling temporal and spectrotemporal regions of interest
- A novel signal processing algorithm for tracking the pitch (fundamental frequency) in noisy bioacoustic recordings
- Dynamic-time-warping and cross-correlation algorithms for comparing batches of vocalizations against each other using pitch or spectrum.
Hosted on GitHub: melizalab/chirp (website)
Publications:
Keen S, Meliza CD, Pilowsky J, Rubenstein DR
(2016). Song in a Social and Sexual Context: Vocalizations Signal Identity and Rank in Both Sexes of a Cooperative Breeder.
Front Ecol Evol,
doi:10.3389/fevo.2016.00046
Keen SC, Meliza CD, Rubenstein DR (2013). Flight calls signal group and individual identity but not kinship in a cooperatively breeding bird. Behav Ecol, doi:10.1093/beheco/art062
Meliza CD, Keen SC, Rubenstein DR (2013). Pitch- and spectral-based time warping methods for comparing field recordings of harmonic avian vocalizations. JASA, doi:10.1121/1.4812269
libtfr
Time-frequency reassignment is a technique for sharpening spectrographic representations of sounds (see figure). For relatively simple, non-stationary processes, TFR can provide substantial improvements in detecting fine structure. Further improvements can be realized by using multiple windowing functions (similar to the multitaper method for calculating the spectra of stationary processes), but the algorithms are computationally intensive. libtfr is a library for calculating multitaper TFR spectrograms which is implemented in C and uses the highly efficient FFTW library. It also supports calculating standard multitaper spectrograms and spectra, and comes with python/numpy and MATLAB interfaces.
Hosted on GitHub: melizalab/libtfr (website)
znote
Vocalizations often consist of spectrotemporally disjoint, discrete elements, and it's often desirable to extract and manipulate these elements separately. However, because they often overlap temporally, the separation can only be achieved in the spectrotemporal domain. znote is a software package for identifying components in bioacoustic signals and extracting the sound pressure waveforms associated with them.
Hosted on GitHub: melizalab/znote
Publications:
Meliza CD, Chi Z, Margoliash D
(2010). Representations of Conspecific Song by Starling Secondary
Forebrain Auditory Neurons: Towards a Hierarchical Framework. J
Neurophys,
doi:10.1152/jn.00464.2009
Data Management
neurobank
The goals of our data-management system are to ensure that the data we collect is stored in a central location where it can be easily located, avoiding needless duplication and assuring that regular backups are taken.
The approach is simple: every resource gets a unique identifier and gets moved to an archive on our main Linux server. Each resource's identifier, archive, and any metadata are are stored in a database with a REST API. There's a commandline tool for depositing resources and for searching and retrieving records from the database.
There are two components to the neurobank system. One part is the registry, a service that stores identifiers, ensures they're unique, and resolves identifiers to the location where the resource is stored. This is implemented by melizalab/django-neurobank.
The second component is an archive, a mechanism for storage and retrieval of resources. This is implemented for locally-accessible filesystems in the melizalab/neurobank package.
Lab tools
bird-colony
A Django application used to manage bird colonies (including breeding colonies). You may find that it can also be used for non-avian species. Useful for tracking bands, pedigree relationships, animal locations, and various events. There's also support for storing information about samples associated with animals in the colony, like genomic DNA or song recordings. Records are searchable and there's a JSON API.
Hosted on GitHub: melizalab/django-bird-colony
lab-inventory
A Django application used to track inventory and orders in the lab. Pretty basic but highly useful. Create items corresponding to specific purchaseable items, then create an order and link items to it when you make a purchase.
Hosted on GitHub: melizalab/django-lab-inventory